Genes play a crucial role in determining many aspects of our biology, from simple traits like eye color to complex diseases like diabetes and cancer. Genome-Wide Association Studies (GWAS) are designed to understand how specific genes, and in particular, their variants, influence phenotypic outcomes.
What is GWAS?
GWAS is a method used to identify genetic markers associated with specific traits or health conditions. By analyzing the genomes of different individuals, researchers can find DNA mutations, typically single nucleotide polymorphisms (SNPs), that are associated with a given phenotype. To do so, researchers compile vast genetic datasets from thousands or even millions of individuals cohorts and record corresponding trait or disease information. From this point, it is a colossal statistical undertaking to determine whether specific SNPs are prevalent among individuals presenting the studied phenotype. You can think of it as finding a needle in a genetic haystack.
Regression analysis is used to quantify the association between a genetic variant and an observed phenotype. The phenotype is normally either a binary outcome (i.e. does this person have the disease?) or a quantitative measure, like someone’s height or weight. Since we also know each individual’s genotype (which is either homozygous or heterozygous with respect to the variant, i.e. ALT/ALT, WT/WT, or ALT/WT), we can use this as the ‘X’ variable and regress against phenotype (‘Y’). For binary outcomes, logistic regression is used instead of linear regression. These regressions will also account for potential confounding variables like age, sex, and diet, which may additionally influence phenotype.
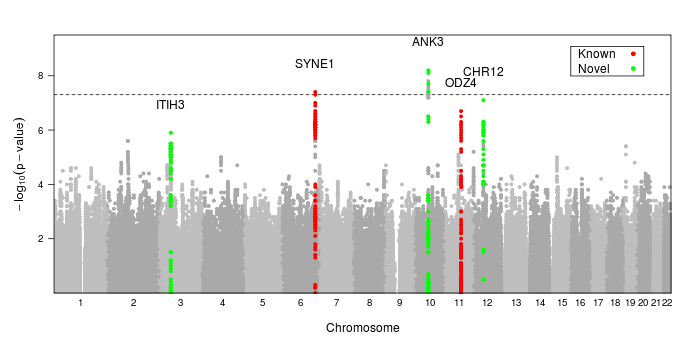
After performing millions of regressions, we end up with a p-value for each variant, indicating the significance of its association to the studied phenotype. These p-values are then corrected for multiple testing, and a cut-off is set to identify which SNPs are significantly associated. Usually, these results are organized into a Manhattan plot (shown below), where p-values are plotted against the genomic positions of the variants they correspond to. Notably, due to a phenomenon called linkage disequilibrium, nearby genetic variants are more likely to be inherited together when DNA is passed down from the parents, so multiple SNPs can end up with lower p-values due to their proximity to a true causal variant. Thus, instead of seeing a single SNP at a higher-than-baseline significance, we often see several SNPs which form the “towers” that you see in the Manhattan plot.
Exploring GWAS on your own
For those interested in the practical side of GWAS, PLINK is a widely used tool that facilitates these analyses. For those not familiar with command-line tools or coding, web-based platforms like GWAS Catalog and SNPedia offer user-friendly interfaces to explore existing GWAS findings.
Clinical and public health applications of GWAS
By revealing the genetic foundations of complex diseases and traits, GWAS can inform initiatives in personalized medicine, where healthcare is customized based on an individual’s genetic profile. GWAS has led to breakthroughs in understanding the genes underlying numerous conditions:
- Heart disease: Variants in the 9p21 region have been identified to increase the risk of coronary artery disease.
- Diabetes: Certain SNPs near the HHEX and CDKN2A/B genes have been linked with type 2 diabetes.
- Macular degeneration: Variants in the CFH gene, and the region on chromosome 10q26, have been associated with age-related macular degeneration.
- Crohn’s disease: Multiple genes, including NOD2 and ATG16L1, have shown a connection to Crohn’s disease.
By identifying several important (i.e. significantly associated genes), GWAS can be used to calculate polygenic risk scores, which predicts an individual’s likelihood to develop a specific disease based on the genotype of several variants. This allows for the stratification of people by their disease risk and facilitates the implementation of preventative strategies for particularly high-risk individuals. We may also look at the prevalence of disease-associated genetic variants in a population to inform public health initiatives. Furthermore, GWAS results can be used to better understand the basis of disease and identify potential targets for drug development, leading to more effective treatments.
Caveats and limitations
While GWAS offers profound insights into phenotype, it is essential to recognize its limitations. GWAS identifies associations rather than causation, and most genetic variants have a relatively small effect on overall risk. Many traits, especially complex diseases, are polygenic and therefore influenced by several genes. The impact of multiple genetic variants on phenotype is not necessarily linear either. Effects due to SNPs, for example, can interact with each other to variably affect phenotype, ultimately producing a non-linear response.
Environmental factors also play a critical role in the development of disease. While GWAS uncovers genetic predispositions, it doesn’t always account for environment influences like diet, exposure to carcinogenic compounds, and exercise.
Moreover, GWAS findings are not always universally applicable across different ancestries, highlighting the need for more diverse and representative samples for research. Historically, GWAS participants have predominantly been of European descent, so the findings from the studies may not generalize well to all ethnic groups. Thus, the application of GWAS to other cohorts is an active area of research.
Conclusion
As GWAS continues to evolve, the insights gleaned from these studies offer us a clearer understanding of our genetic infrastructure. GWAS is a powerful tool, but it is not without its limitations. Nonetheless, this knowledge brings us closer to designing preemptive health strategies and treatments attuned to the individual rather than the multitude.
